Spatkin
From Laboratory of Modeling in Biology and Medicine
A simulator for rule-based modeling on surfaces
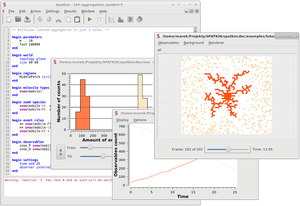
Spatkin implements a direct, lattice-based method, which avoids enumeration of the reaction network implied by a set of rules. Models are specified using an extension of the BioNetGen language, which adds features necessary for characterizing diffusion properties of biomolecules. The tool is capable of simulating biomolecular interactions occurring on or at a two-dimensional surface, such as interactions of the kind commonly involved in receptor signaling.
Downloads
Spatkin is distributed under GNU GPL v3.0. The latest Spatkin version is 1.0.0.
Source code
- C++ source code: spatkin-1.0.0-source.tar.gz
- Compilation instructions
Compilation on Linux, Mac, and Windows is managed by CMake – see file COMPILING.txt in the source code for detailed building instructions.
Stand-alone executables
- macOS: application bundle (zipped "app"): spatkin-1.0.0-x86_64-darwin16.zip
- macOS: verification and running
COMPATIBILITY: This software was built with XCode 8 and tested under macOS 10.12 (Sierra). It is likely incompatible with previous Mac operating system releases.
SECURITY: MD5 sum of the downloadable ZIP archive can be checked in Terminal:md5 path_to_zipMD5 sum of untainted spatkin-1.0.0-x86_64-darwin16.zip is b841ced9da16a6cff9ab2f25654c9334
EXAMPLES: Example Spatkin input files can be found inside Spatkin.app, in folder Documentation. Independently, they can be downloaded here.
- Windows 64-bit: installer: spatkin-1.0.0-x86_win64-setup.exe
- Windows 64-bit: verification and running
COMPATIBILITY: This software was built with MSYS2/MinGW64 on Windows 10, and tested on Windows 10 and Windows 8.
SECURITY: The downloadable installer is not signed with a certificate (so in windows terms it comes from an "unknown publisher") but its integrity can be checked manually in command line using a Windows pre-installed utility:certUtil -hashfile path_to_installer MD5MD5 sum of untainted spatkin-1.0.0-x86_win64-setup.exe is:
e3 27 16 0b 5a 0e a1 b7 f8 b8 e7 7b 46 c0 26 ec
EXAMPLES: After installation, example Spatkin input files can be found in the windows user directory Documents/SpatkinModels. (There is a new set of examples; installer asks whether the previously installed set of examples should be removed; it's recommended to remove previous examples.) Independently, the same set of examples can be downloaded here.
Virtual machines with pre-installed Spatkin
- Virtual machine hosting Ubuntu Linux: Virtual_machine_with_spatkin.7z
- Setup instructions
The 7-zip archive contains a configured virtual machine that was created under VirtualBox 5.1.22. Virtual disk format is VMDK; the guest operating system installed on the disk should thus run not only under VirtualBox but also under VMWare, QEMU, and other virtualization solutions. The machine hosts Ubuntu Linux 16.04.2 LTS.
PLATFORM: VirtualBox for the major operating systems (as hosts) can be downloaded free of charge from http://www.virtualbox.org/wiki/Downloads.
LAUNCHING: After downloading, the archive should be decompressed (command-line: 7z x Virtual_machine_with_spatkin.7z). In Oracle VirtualBox Manager, one should choose from the menu Machine -> Add and point to the SpatkinLinuxBox.vbox file. In the guest Linux system, the default user is ubuntu (password: ubuntu); this user is automatically logged after startup. Spatkin can be launched via a desktop shortcut or by opening terminal and typing spatkin.
EXAMPLES: Example SPATKIN input files can be found in /home/ubuntu/local/spatkin/share/doc. Independently, they can be retrieved, most easily, using wget in Terminal:
wget https://pmbm.ippt.pan.pl/software/spatkin/release/spatkin-1.0.0-examples.zip
POSSIBLE ISSUES: An issue that often arises when launching a virtual machine within VirtualBox for the first time is the lack of hardware-accelerated virtualization technologies such as VT-x or AMD-V. Please note that you may need to enable this feature by changing advanced CPU settings in BIOS of your computer (support for virtualization is often disabled by default).
[new] Containers
- Dockerfile to build a Docker container image: spatkin-1.0.0-Dockerfile
- A docker-based recipe for a singularity container: spatkin-1.0.0-singularity.recipe
- Setup instructions
BUILDING a Singularity container (in terminal):$ singularity build spatkin.simg spatkin-1.0.0-singularity.recipe
LAUNCHING the simulator within the sinfularity container (in terminal):export LC_ALL=C
$ singularity exec spatkin.simg /opt/local/bin/Spatkin
BUILDING a Docker container (in terminal):$ docker build --build-arg user=$USER --build-arg uid=$(id -u) --build-arg gid=$(id -g) -t spatkin .
RUNNING the Docker container (in terminal):$ xauth nlist $DISPLAY | sed -e 's/^..../ffff/' | xauth -f /tmp/.docker.xauth nmerge -
$ docker run -e DISPLAY=unix$DISPLAY -v /tmp/.X11-unix:/tmp/.X11-unix -v /tmp/.docker.xauth:/tmp/.docker.xauth:rw -e XAUTHORITY=/tmp/.docker.xauth -v $HOME:$HOME spatkin
Documentation
- User manual (contains annotated example input files): spatkin-1.0.0-manual.pdf
Example models
- An archive with example Spatkin files: spatkin-1.0.0-examples.zip
Contact
The lead software developer is Marek Kochańczyk. For both user's and developer's queries please use the following e-mail address: spatkin.simulator@gmail.com
References
Primary software citation
- Kochańczyk M, Hlavacek WS, Lipniacki T. Spatkin: a simulator for rule-based modeling of biomolecular site dynamics on surfaces, Bioinformatics (2017) 33(22):3667–3669 PubMed CrossRef
Spatkin simulation-based studies
- Nałęcz-Jawecki P, Szymańska P, Kochańczyk M, Miękisz J, Lipniacki T. Effective reaction rates for diffusion-limited reaction cycles, J Chem Phys 143(21):215102 (2015) PubMed CrossRef | PDF-ms
- Szymańska P☯, Kochańczyk M☯, Miększ J, Lipniacki T. Effective reaction rates in diffusion-limited phosphorylation–dephosphorylation cycles, Phys Rev E 91:022702 (2015) PubMed CrossRef | PDF-ms
- Kochańczyk M, Jaruszewicz J, Lipniacki T. Stochastic transitions in a bistable reaction system on the membrane, J R Soc Interface 10(84):20130151 (2013) PubMed CrossRef | PDF Supp-PDF Supp-movies
- Żuk PJ, Kochańczyk M, Jaruszewicz J, Bednorz W, Lipniacki T. Dynamics of a stochastic spatially extended system predicted by comparing deterministic and stochastic attractors of the corresponding birth-death process, Phys Biol 5(9):055002 (2012) PubMed CrossRef | PDF-ms